Bioinformatic Support
Bioinformatic Support
Bioinformatics is now essential to modern biological research.
By combining biology, computer science, and statistics, we provide end-to-end bioinformatics solutions for projects of any size. We offer training, ongoing support, and bioinformatics-as-a-service, tailored to your project’s needs.
Overview
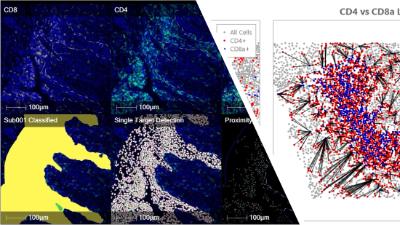
Our multiplex immunofluorescence platforms (Cell DIVE and PhenoCycler) generate exceptionally rich datasets—often hundreds of gigabytes from a single slide.
Using HALO digital pathology software with the HiPlex FL module, we extract quantitative biomarker intensity and spatial positioning data for every individual cell within imaged tissues.
We support multiplex image analysis across the full spectrum, from straightforward quantitative measurements to advanced, high-resolution spatial analyses.
Example Analysis Types
-
Area-based biomarker quantification
Measurement of tissue area positive for specific markers (e.g. ECM components) -
Cell type enumeration
Quantification of cell populations within defined tissue regions or layers -
Unbiased cell phenotyping
Data-driven classification of cellular phenotypes -
Cell neighbourhood analysis
Characterisation of local cellular microenvironments -
Spatial analysis
Distance metrics, attraction/avoidance patterns, and immune infiltration analysis
Overview
Our high-content imaging platforms (INCA2200 and INCA6000) combined with dedicated analysis software (INCarta and Developer) enable fully automated analysis of slides and multi-well plates.
We have extensive experience working with high-content imaging data to deliver high-quality, publication-ready analyses—ranging from straightforward quantitative readouts to advanced machine learning and AI-driven models.
Example Analysis Types
- Cell-level quantification
Simple cell counts and population statistics -
Marker-positive analysis
Thresholding and percentage calculations -
Organelle-level analysis
Subcellular features such as DNA damage foci -
Large-scale morphological profiling
Including senescence-associated morphological profiles (SAMPs) -
Advanced visualisation and dimensionality reduction
Heatmaps, frequency distributions, and embeddings (UMAP, t-SNE)
Overview
A single high-throughput screen can generate thousands of images across multi-well plates, often spread across multiple batches, cell types, and treatment groups.
We have extensive experience handling large-scale screening datasets, supporting robust and reproducible readouts ranging from hit identification to rare-event discovery and batch normalisation.
Example Analysis Types
- Hit identification and ranking
Z-scores and other statistical ranking methods - Phenocopying
Identification of treatments or perturbations producing similar phenotypic profiles - Rare event detection
Discovery of low-frequency but biologically relevant phenotypes - Batch correction and normalisation
Minimising technical variation across plates, runs, and experiments - Quality control and assay performance metrics
Plate effects, control performance, and reproducibility assessment - Data mining of archival screens
Reanalysis of legacy datasets to uncover new phenotypes, targets, or treatment relationships
We provide training and ongoing support for all dedicated facility software, tailoring analysis pipelines to the specific needs of your project. We can teach you how to perform the analysis yourself, or build and run the complete analysis for you as a service.
We also offer a general introduction to scripting (e.g. R / RStudio), with a focus on how to integrate custom scripts alongside data generated within the facility.
In addition, we maintain a curated library of scripts, tips, and best practices designed to accelerate analysis, improve reproducibility, and help you get results faster.
Bioinformatics support is available to both internal and external users.
-
Internal users: £35 per hour
-
External users: £55 per hour
We strongly recommend including bioinformatics support time in project grant applications, allowing us to provide the most effective and timely support for your project.
For further information or to book time, please contact our Lead Bioinformatician Dr Ryan Wallis:
phenotypic-screening@qmul.ac.uk